MiPNet21.06 SUIT RP: Difference between revisions
No edit summary |
|||
| Line 22: | Line 22: | ||
== SUIT reference protocol-RP1 == | == SUIT reference protocol-RP1 == | ||
[[File: | [[File:1PM;2D;2c;3U;4G;5S;6Oct;7Rot;8Gp-9Ama;10Tm;11Azd.jpg|right|700px|link=SUIT-RP1 |SUIT-RP1]]<br /> | ||
****:» [[SUIT_FNSGp(PGM)01]] | ****:» [[SUIT_FNSGp(PGM)01]] | ||
****:» [[SUIT_FNSGp(PGM)02]] | ****:» [[SUIT_FNSGp(PGM)02]] | ||
Revision as of 11:46, 14 March 2017
| SUIT reference protocol for OXPHOS analysis by high-resolution respirometry. |
OROBOROS (2016-08-17) Mitochondr Physiol Network
Abstract: Doerrier C, Sumbalova Z, Krumschnabel G, Hiller E, Gnaiger (2016) SUIT reference protocol for OXPHOS analysis by high-resolution respirometry. Mitochondr Physiol Network 21.06(01):1-12. » Versions
- » MitoPedia: SUIT reference protocol
- » Instrument: O2k, O2k-Catalogue
• O2k-Network Lab: AT_Innsbruck_OROBOROS
Labels: MiParea: Instruments;methods
Preparation: Permeabilized cells, Permeabilized tissue, Homogenate, Isolated mitochondria
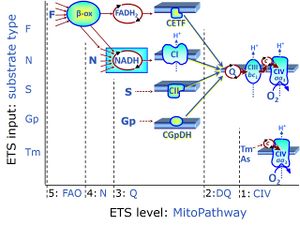
Regulation: Substrate Coupling state: LEAK, OXPHOS, ETS"ETS" is not in the list (LEAK, ROUTINE, OXPHOS, ET) of allowed values for the "Coupling states" property. Pathway: F, N, S, Gp, CIV, NS, Other combinations HRR: Oxygraph-2k, O2k-Protocol
MitoPathways, MitoFitPublication, 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp-, 1D;2OctM;3P;4G;5S;6Gp;7U;8Rot-
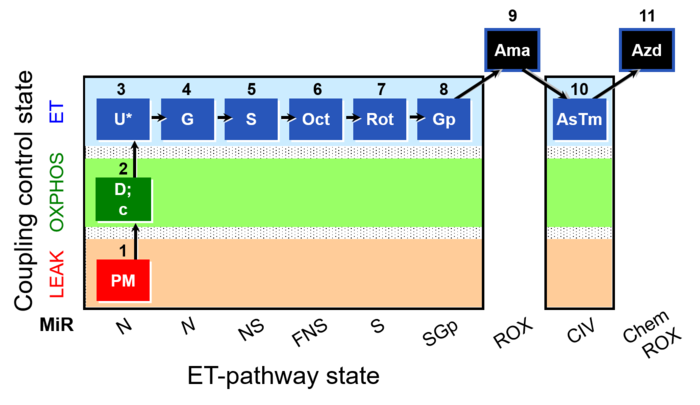
SUIT reference protocol-RP1
SUIT states: 1-4PM(LPcE) 5PGM 6PGMS 7OctPGMS 8S 9SGp 10ROX 11Tm 12ROX
| Step | Respiratory state | Pathway control | Pathway to Q |
|---|---|---|---|
| 1PM | PM(L) | N | CI |
| 2D | PM(P) | N | CI |
| 3c | PM(P)c | N | CI |
| 3NADH | PM(P)cNADH | N | CI |
| 4U | PM(E) | N | CI |
| 5G | PGM(E) | N | CI |
| 6S | PGMS(E) | NS | CI&II |
| 7Oct | OctPGMS(E) | FNS | FAO&CI&II |
| 8Rot | S(E) | S | CII |
| 9Gp | SGp(E) | SGp | CII&GpDH |
| 10Ama | ROX | ROX | |
| 11Tm | Tm(E) | CIV | CIV |
| 12Azd | ROX | ROX |
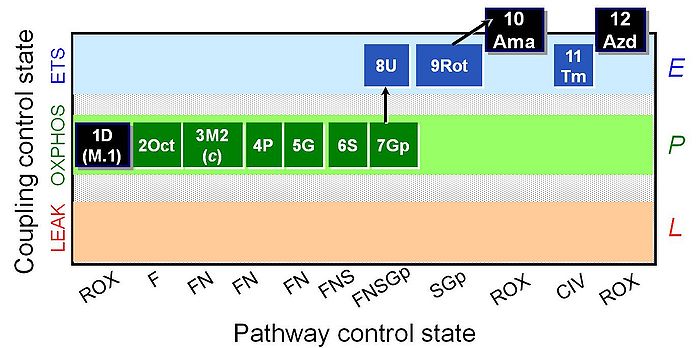
SUIT reference protocol-RP2
SUIT states: 1ROX 2OctM(P) 3OctM(c) 4OctPM 5OctPGM 6OctPGMS 7OctPGMSGp 8E 9SGp 10ROX 11CIV 12ROX
| Step | Respiratory state | Pathway control | Pathway to Q |
|---|---|---|---|
| 1D | ROX | ROX | |
| (M.1) | |||
| 2Oct | Oct(P) | (F) | FAO |
| 3M2 | OctM(P) | F | FAO |
| (c) | OctM(P)c | F | FAO |
| 4P | OctPM(P) | FN | FAO&CI |
| 5G | OctPGM(P) | FN | FAO&CI |
| 6S | OctPGMS(P) | FNS | FAO&CI&II |
| 7Gp | OctPGMSGp(P) | FNSGp | FAO&CI&II&GpDH |
| 8U | OctPGMSGp(E) | FNSGp | FAO&CI&II&GpDH |
| 9Rot | SGp(E) | SGp | CII&GpDH |
| 10Ama | ROX | ROX | |
| 11Tm | Tm(E) | CIV | CIV |
| 12Azd | ROX | ROX |
Laboratory sheets for SUIT protocols RP1 and RP2
»RP1 for mitochondrial preparations (mt), permeabilized cells (pce) and permeabilized fibers (pfi)
»RP2 for mitochondrial preparations (mt), permeabilized cells (pce) and permeabilized fibers (pfi)
DatLab-Analysis templates for RP analysis (DatLab 7)
DatLab-Analysis templates for RP analysis (respiration and fluorescence: H2O2 production)
Experimental details
Work in progress: pre-publication protocol
Saturating ADP concentrations
- Different concentrations of ADP are sufficient to obtain maximum flux for estimating OXPHOS capacity. The typical range for isolated mitochondria is 1 to 2.5 mM, for permeabilized cells 1 to 5 mM, for permeabilized muscle fibres 2.5 to 10 mM.
RP-T01
- RP1: Compared to RP1-T01, move Oct titration after S.
- RP2: Compared to RP2-T01, move U titration after Gp.
RP-T02
- RP2: Compared to RP2-T02, move D titration after the sample. Titration of M.1 before Oct. M2 is added after Oct.
Cytochrome c test
- In protocol RP2, the cytochrome c test might better be moved from a c-titration after pyruvate (P) to a titration after M2. Otherwise, there is a risk of underestimation of FAO in cases of a cytochrome c effect. On the other hand, at the low flux before titration of P, the sensitivity of the cytochrome c test may be low.
- ~ Gnaiger Erich 18:26, 23 January 2016 (CET)
Mark names
- Even the highly abbreviated names of respiratory states have become too long for DatLab, where mark names are restricted to 8 digits. The visibility in DatLab of long mark names is restricted. The simplest solution for short mark names is the use of a numerical sequence, with the preceeding event name added for information.
- ~ Gnaiger Erich 16:50, 23 January 2016 (CET)
Pre-publication communication
- This communication is a pre-publication, inviting critical feedback and comments. All feedback will be carefully documented and evaluated in terms of justification of co-authorship. We intend to finally publish this topic in a peer-reviewed journal as an original article, with reference to the pre-publication history including reviewer's reports. - Gnaiger Erich 03:23, 18 January 2016 (CET)
Experiments in progress
- The SUIT reference protocol is presently applied in permeabilized HEK cells, mouse heart isolated mitochondria, liver homogenate, permeabilized skeletal muscle (mouse and human), and human PBMCs and platelets. - Gnaiger Erich 18:47, 19 January 2016 (CET)
- AT Innsbruck Gnaiger E and AT Innsbruck OROBOROS - MitoFit project.
- US NC Winston-Salem Molina AJA - UPBEAT project.
Further details
- » Introduction: Gnaiger 2014 MitoPathways
- » MitoPedia: SUIT - extended MitoPedia topic, May 2016
- Table of titrations » MiPNet09.12 O2k-Titrations
- » Definition: Substrate-uncoupler-inhibitor titration
- » Context: SUIT protocol library
- » Abbreviations: MitoPedia