SUIT-001: Difference between revisions
From Bioblast
No edit summary |
Timon Alba (talk | contribs) No edit summary |
||
| (9 intermediate revisions by 3 users not shown) | |||
| Line 12: | Line 12: | ||
{{MitoPedia O2k and high-resolution respirometry}} | {{MitoPedia O2k and high-resolution respirometry}} | ||
{{MitoPedia topics}} | {{MitoPedia topics}} | ||
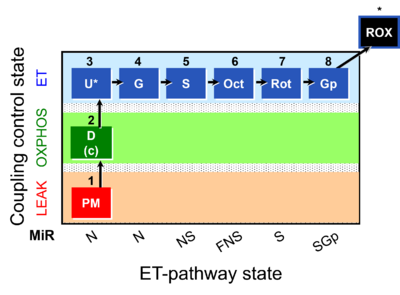
::'''[[SUIT protocol pattern]]''': 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp- | |||
The SUIT-001 protocol in combination with [[SUIT-002]] is specially designed to provide a common reference for comparison of respiratory control of [[Mitochondrial preparations| mitochondrial preparations]] in a wide variety of species, tissues and cell types. SUIT-001 gives information of the linear coupling control ([[LEAK | The SUIT-001 protocol in combination with [[SUIT-002]] is specially designed to provide a common reference for comparison of respiratory control of [[Mitochondrial preparations| mitochondrial preparations]] in a wide variety of species, tissues and cell types. SUIT-001 gives information of the linear coupling control ([[LEAK respiration|''L'']]-[[Oxidative phosphorylation| ''P'']]-[[ET capacity| ''E'']]) with NADH linked-substrates ([[PM-pathway control state|PM]]). Moreover, the pathway control in ET state ([[NADH_Electron_transfer-pathway_state|N]], [[NS-pathway control state| NS]], [[FNS]], [[Succinate pathway control state| S]] and [[SGp-pathway control state| SGp]] pathways) can be evaluated by using this SUIT protocol. SUIT-001 can be extended with the CIV assay module. | ||
__TOC__ | __TOC__ | ||
| Line 20: | Line 20: | ||
== Specific SUIT protocols == | == Specific SUIT protocols == | ||
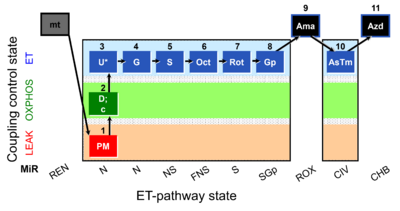
=== SUIT-001 O2 mt D001 | === SUIT-001 O2 mt D001 === | ||
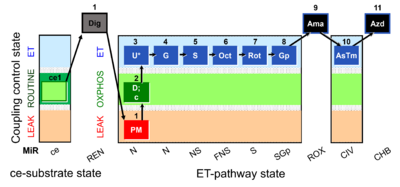
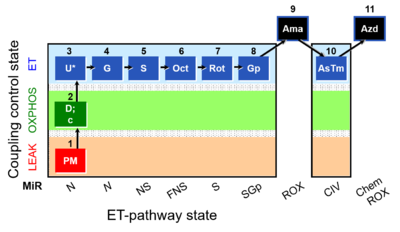
[[File:1PM;2D;2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10AsTm;11Azd.png|400px]] | [[File:Mt;1PM;2D;2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10AsTm;11Azd .png|400px]] | ||
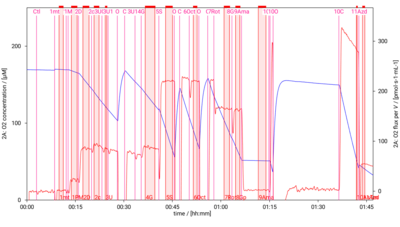
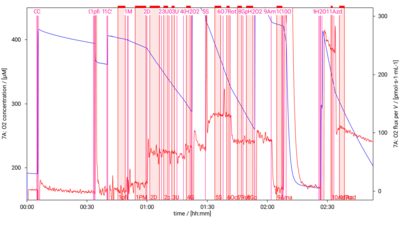
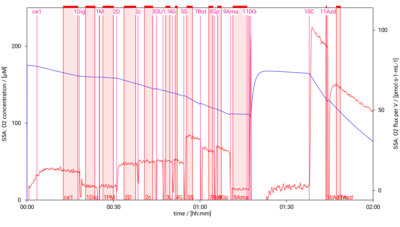
[[File:D001_O2_traces.png|400px]] | [[File:D001_O2_traces.png|400px]] | ||
* [[SUIT-001 O2 mt D001]] for [[Isolated mitochondria|isolated mitochondria]], [[Tissue homogenate|tissue homogenate]] and [[Permeabilized cells|permeabilized cells]] (already permeabilized when they are added to the chamber) | * [[SUIT-001 O2 mt D001]] for [[Isolated mitochondria|isolated mitochondria]], [[Tissue homogenate|tissue homogenate]] and [[Permeabilized cells|permeabilized cells]] (already permeabilized when they are added to the chamber) | ||
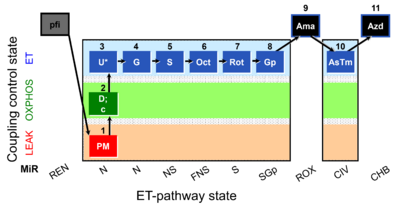
=== SUIT-001 O2 pfi D002 === | |||
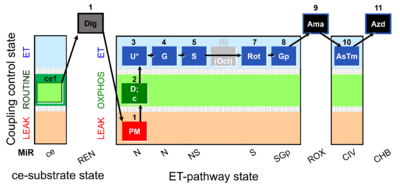
[[File:Pfi;1PM;2D;2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10AsTm;11Azd.png|400px]] | |||
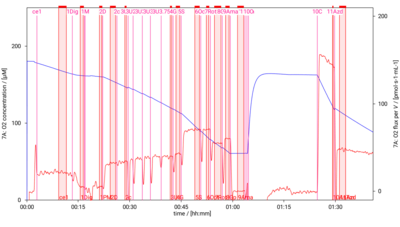
[[File:D002 O2 traces.png|400px]] | |||
* [[SUIT-001 O2 pfi D002]] for [[Permeabilized_muscle_fibers|permeabilized muscle fibers]] | * [[SUIT-001 O2 pfi D002]] for [[Permeabilized_muscle_fibers|permeabilized muscle fibers]] | ||
| Line 34: | Line 38: | ||
[[File:ce1;1Dig;1PM;2D;2c;3U;4G;5S;(6Oct);7Rot;8Gp;9Ama;10AsTm;11Azd.png|400px]] | [[File:ce1;1Dig;1PM;2D;2c;3U;4G;5S;(6Oct);7Rot;8Gp;9Ama;10AsTm;11Azd.png|400px]] | ||
[[File:D004_O2_traces.png|400px]] | [[File:D004_O2_traces.png|400px]] | ||
* [[SUIT-001 O2 ce-pce D004]] for [[Living cells|living cells]] (ce) and [[Permeabilized cells|permeabilized | * [[SUIT-001 O2 ce-pce D004]] for [[Living cells|living cells]] (ce) and [[Permeabilized cells|permeabilized PBMC and PLT]] (pce; permeabilization of the plasma membrane occurs into the chambers through digitonin addition)(SUIT-001 O2 ce-pce D004 is performed in the absence of Oct). | ||
| Line 57: | Line 61: | ||
:::- The substrate combination for maximum ET capacity (FNSGp) is not obtained in SUIT-001 in favour of measuring S_E. | :::- The substrate combination for maximum ET capacity (FNSGp) is not obtained in SUIT-001 in favour of measuring S_E. | ||
:::- Oct is not added in | :::- Oct is not added in PBMC and PLT ([[SUIT-001 O2 ce-pce D004]]). | ||
:::- Lengthy duration of the experiment. | :::- Lengthy duration of the experiment. | ||
== Compare SUIT protocols == | == Compare SUIT protocols == | ||
::::* [[SUIT-001 O2 ce-pce D004]] (SUIT-001 for permeabilized | ::::* [[SUIT-001 O2 ce-pce D004]] (SUIT-001 for permeabilized PBMC and PLT) differs from [[SUIT-001 O2 ce-pce D003]] (general SUIT-001 for permeabilized cells) in the absence and presence of octanoylcarnitine, respectively. | ||
::::* [[SUIT-002]] (RP2): Harmonized SUIT protocol. | ::::* [[SUIT-002]] (RP2): Harmonized SUIT protocol. | ||
Latest revision as of 16:38, 5 March 2024
Description
Abbreviation: RP1
Reference: Doerrier 2018 Methods Mol Biol
MitoPedia concepts:
MiP concept,
SUIT protocol,
Recommended
MitoPedia methods:
Respirometry
- SUIT protocol pattern: 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp-
The SUIT-001 protocol in combination with SUIT-002 is specially designed to provide a common reference for comparison of respiratory control of mitochondrial preparations in a wide variety of species, tissues and cell types. SUIT-001 gives information of the linear coupling control (L- P- E) with NADH linked-substrates (PM). Moreover, the pathway control in ET state (N, NS, FNS, S and SGp pathways) can be evaluated by using this SUIT protocol. SUIT-001 can be extended with the CIV assay module.
Communicated by Doerrier C, Gnaiger E (last update 2019-06-07)
Specific SUIT protocols
SUIT-001 O2 mt D001
- SUIT-001 O2 mt D001 for isolated mitochondria, tissue homogenate and permeabilized cells (already permeabilized when they are added to the chamber)
SUIT-001 O2 pfi D002
SUIT-001 O2 ce-pce D003
- SUIT-001 O2 ce-pce D003 for living cells (ce) and permeabilized cells (pce; permeabilization of the plasma membrane occurs into the chambers through digitonin addition)
SUIT-001 O2 ce-pce D004
- SUIT-001 O2 ce-pce D004 for living cells (ce) and permeabilized PBMC and PLT (pce; permeabilization of the plasma membrane occurs into the chambers through digitonin addition)(SUIT-001 O2 ce-pce D004 is performed in the absence of Oct).
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1PM | PML(n) | N | CI | 1PM
|
| 2D | PMP | N | CI | 1PM;2D
|
| 2c | PMcP | N | CI | 1PM;2D;2c
|
| 3U | PME | N | CI | 1PM;2D;3U
|
| 4G | PGME | N | CI | 1PM;2D;3U;4G
|
| 5S | PGMSE | NS | CI&II | 1PM;2D;3U;4G;5S
|
| 6Oct | OctPGMSE | FNS | FAO&CI&II | 1PM;2D;3U;4G;5S;6Oct
|
| 7Rot | SE | S | CII | 1PM;2D;3U;4G;5S;6Oct;7Rot
|
| 8Gp | SGpE | SGp | CII&GpDH | 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp
|
| 9Ama | ROX | 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp;9Ama
|
| Step | Respiratory state | Pathway control | ET-Complex | Comment |
|---|---|---|---|---|
| ## AsTm | AsTmE | CIV | CIV | |
| ## Azd | CHB |
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- SUIT-001 in combination with SUIT-002 provides a common reference for comparison of respiratory control in a large variety of species, tissues and cell types. Both SUIT protocols provide a mitochondrial mapping which allows:
- 1. to obtain reference values.
- 2. to evaluate mitochondrial physiological diversity, generating a mt-database on comparative mitochondrial physiology.
- 3. to screen specific defects.
- SUIT-001 and SUIT-002 are used in the MitoFit Proficiency test for inter-individual and inter-laboratory reproducibility quality control.
- A succinate concentration of >10 mM may be required for saturating S_E capacity.
- + Linear coupling control (L-P-E) with N substrates (PM).
- + Pathway control in ET state (N, NS, FNS, S and SGp pathways).
- + Harmonization with SUIT-002 allows to perform both SUIT protocols in parallel. The cross-linked respiratory states can be statistically used as repeated measurements.
- + Harmonization with many SUIT protocols (up to step 5S).
- + Oct is added in ET state to avoid uncoupler effect with fatty acids.
- + If NS- and FNS-pathways are similar (NS_E ~ FNS_E), the additivity index of N- and S-pathways in ET state can be evaluated.
- + This protocol can be extended with the Complex IV module.
- - The substrate combination for maximum ET capacity (FNSGp) is not obtained in SUIT-001 in favour of measuring S_E.
- - Oct is not added in PBMC and PLT (SUIT-001 O2 ce-pce D004).
- - Lengthy duration of the experiment.
Compare SUIT protocols
- SUIT-001 O2 ce-pce D004 (SUIT-001 for permeabilized PBMC and PLT) differs from SUIT-001 O2 ce-pce D003 (general SUIT-001 for permeabilized cells) in the absence and presence of octanoylcarnitine, respectively.
- SUIT-002 (RP2): Harmonized SUIT protocol.
References
| Year | Reference | Organism | Tissue;cell | |
|---|---|---|---|---|
| Doerrier 2018 Methods Mol Biol | 2018 | Doerrier C, Garcia-Souza LF, Krumschnabel G, Wohlfarter Y, Mészáros AT, Gnaiger E (2018) High-Resolution FluoRespirometry and OXPHOS protocols for human cells, permeabilized fibers from small biopsies of muscle, and isolated mitochondria. Methods Mol Biol 1782:31-70. https://doi.org/10.1007/978-1-4939-7831-1_3 | Human Mouse Rat Saccharomyces cerevisiae | Heart Skeletal muscle Endothelial;epithelial;mesothelial cell Blood cells HEK Platelet |