Description
Abbreviation: NS(GM)
Reference: A:short protocol for simultaneous determination of O2 flux and mitochondrial membrane potential in mitochondrial preparations (isolated mitochondria, permabilized cells, and tissue homogenate)-SUIT-021
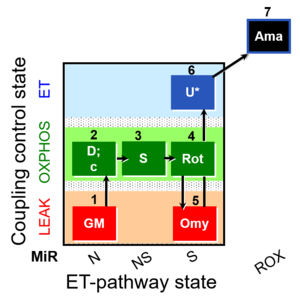
SUIT number: D035_1GM;2D;2c;3S;4Rot;5Omy;6U;7Ama
O2k-Application: O2
- SUIT-category: NS(GM)
- SUIT protocol pattern: orthogonal 1GM;2D;2c;3S;4Rot;5Omy;6U-
SUIT-021 O2 mt D035 is design to assess the additivity between N-pathway and S- pathway in the Q-junction and to investigate the N- and NS-pathway control state in mitochondrial preparations . Oligomycin (Omy) is used to induce a LEAK state of respiration through inhibition of the ATP synthase. Higher concentration of Omy can decreases the ET state initiated by an uncoupler, therefore the required concentration of Omy has to be determined. Uncoupler increases the respiration and induces the ET state. Antimycin A (Ama), which is an inhibitor of Complex III), blocks the respiration. Originially this is a protocol for simultaneous determination of O2 flux and mitochondrial membrane potential on isolated mitochondria and tissue homogenate. It can be used as a control experiment for SUIT-021 Fluo mt D036 without using the fluorescence dye in order to evaluate its effect on the O2 flux as a control.
Communicated by Huete-Ortega M, Komlodi T and Gnaiger E (last update 2019-04-03)
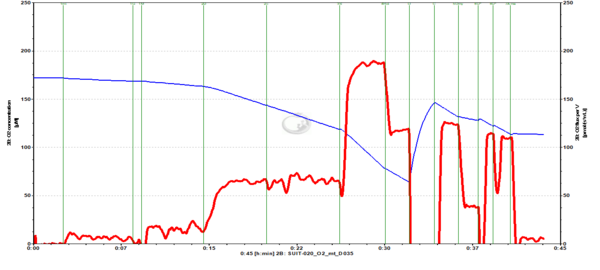
Representative traces
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1GM | GML(n) | N | CI | 1GM
|
| 2D | GMP | N | CI | 1GM;2D
|
| 2c | GMcP | N | CI | 1GM;2D;2c
|
| 3S | GMSP | NS | CI&II | 1GM;2D;2c;3S
|
| 4Rot | SP | S | CII | 1GM;2D;2c;3S;4Rot
|
| 5Omy | SL(Omy) | 1GM;2D;2c;3S;4Rot;5Omy
| ||
| 6U* | SE | S | CII | 1GM;2D;2c;3S;4Rot;5Omy;6U
|
| 7Ama | ROX | 1GM;2D;2c;3S;4Rot;5Omy;6U;7Ama
|
| Step | Respiratory state | Pathway control | ET-Complex | Comment |
|---|---|---|---|---|
| ## AsTm | AsTmE | CIV | CIV | |
| ## Azd | CHB |
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- NS-OXPHOS capacity provides a physiologically relevant estimate of maximum mitochondrial respiratory capacity.
- Comparison of GM- with PM-capacity yields important information on N-pathway respiratory control upstream of CI.
- A succinate concentration of >10 mM may be required for saturating SE capacity.
- + You obtain information in a single protocol about the NS-, S and N- pathway control state.
- + CIV activity determination and Cytochrome c test can be performed.
- - Oligomycin concentration has to be determined. Higher concentrations of oligomycin may inhibit the ET state.
- - Careful washing is required after the experiment to avoid carry-over of the inhibitors and uncoupler.
Compare SUIT protocols
- SUIT-001;SUIT-001 O2 mt D001for mitochondrial preparations: a complex protocol to get information not only about the NS(PGM) in the OXPHOS state, but also in ET state and about the FGp-pathways.
- SUIT-004;SUIT-004 O2 pfi D010: provides information about the NS(PM) pathway in the ET state without the contribution of G.
- SUIT-008;SUIT-008 O2 pfi D014 and SUIT-008 O2 ce-pce D025: provide information about the NS(PGM) pathway in the OXPHOS and ET-states, but without the addition of Omy.
- SUIT-011: provides information about the NS(GM) pathway in the OXPHOS and ET-states, but without the addition of Omy.
- SUIT-012: provides information about the N(PGM) pathway in the OXPHOS and ET-states without the contribution of S-pathway and without the addition of Omy.
- SUIT-014: a protocol similar to SUIT-021 O2 mt D035, but with the addition of P in the OXPHOS state before S. Omy is not added in the protocol.
Chemicals and syringes
| Step | Chemical(s) and link(s) | Comments |
|---|---|---|
| 1GM | Glutamate (G) and Malate (M) | |
| 2D | ADP (D) | |
| 2c | Cytochrome c (c) | |
| 3S | Succinate (S) | |
| 4Rot | Rotenone (Rot) | |
| 5Omy | Oligomycin (Omy) | |
| 6U | Carbonyl cyanide m-chlorophenyl hydrazone, CCCP (U) | Can be substituted for other uncoupler |
| 7Ama | Antimycin A (Ama) |
- Suggested stock concentrations are shown in the specific DL-Protocol.
References
MitoPedia concepts: SUIT protocol, SUIT A, Find
MitoPedia methods:
Respirometry