Vella 2019 MiPschool Coimbra
| Genetic variants in mitochondrial respiratory chain complex deficiencies. |
Link: MitoEAGLE
Vella J, Laurie S, Matalonga L, Borg J, Soler D, Said E, Felice A (2019)
Event: MiPschool Coimbra 2019
Mitochondrial disorders are considered are genetically heterogenous. The oxidative phosphorylation (OXPHOS) system consists of four multi-protein enzyme complexes CI, CII, CIII, CIV and ATP synthase [1]. We report here two cases of patients suspected to have a mitochondrial disorder whose samples were banked at the University of Malta’s biobank (UM biobank) [2].
The analysis was part of a collaborative BBMRI-Large Prospective Cohort (BBMRI-LPC) project focused on mitochondrial disorders. The full mitochondrial genome sequenicng, whole exome sequencing (WES) and data processing were carried out at Centro Nacional de Análisis Genómico (CNAG-CRG). Phenotypic data was recorded in the RD-Connect PhenoTips instance, and variant filtration and prioritisation was undertaken using the RD-Connect Genome-Phenome Analysis Platform [3].
Patient A (Pt A) inherited nuclear genetic variants in 4 of the 7 ‘core’ membrane embedded CI subunits it encodes: NDUFS1 (N module); NDUFA10 (in ND2); NDUFB9 (in ND5), and a rare homozygous mis-sense variant c.308C>T (rs749249430) in NDUFAF3, a CI assembly factor (Q module). Pt A is also a carrier for TIMM13 and CLPX (Table 1, Figure 1).
Patient B inherited a mt DNA missense mutation in MT-ATP6 c.163A>G at m.8689 and 3 X-linked splicing variants: c.207+2T>G (rs782792601); c.206A>G (rs781909386); and c.205A>G (rs782503581) in NDUFB11 (in ND4) (Table 1, Figure 1).
Mutations in encoding subunits and assembly factors of CI cause mt CI deficiency and mitochondrial disease. A novel variant in ATP6 (m.8689) is associated with mt complex V deficiency and Leigh syndrome. It is thought that multiple mutations on different loci could be responsible for complex phenotypes. A cell line model is being used to study the molecular mechanisms involved in CI and ATP synthase assembly.
WES followed by functional validation of disease alleles could identify disease-causative variants in mitochondrial respiratory chain complexes.
• Bioblast editor: Plangger M
Labels: MiParea: mtDNA;mt-genetics, nDNA;cell genetics, Patients
Stress:Mitochondrial disease
Enzyme: Complex I
Affiliations and support
- Vella J(1), Laurie S(2), Matalonga L(2), Borg J(1,3), Soler D(4), Said E(4), Felice A(1,3,4)
- Centre Molecular Medicine Biobanking, Univ Malta, Msida, Malta,
- Centro Nacional Análisis Genómico (CNAG-CRG), Center Genomic Regulation, Barcelona Inst Science Technology (BIST), Univ Pompeu Fabra (UPF), Barcelona, Spain
- Dept Applied Biomedical Science, Physiology Biochemistry, Univ Malta, Msida
- Dept Paediatrics Pathology, Mater Dei Hospital; Malta.
- Vella J(1), Laurie S(2), Matalonga L(2), Borg J(1,3), Soler D(4), Said E(4), Felice A(1,3,4)
- The research leading to these results has received funding from the European Community’s Seventh Framework Programme (FP7/2007-2013) under grant agreement no. 2012-305444, the 2016 BBMRI-LPC WES call and the Malta Government Scholarship Scheme.
Figures and Tables
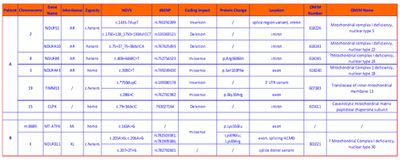
Table 1: Candidate variants from WES.
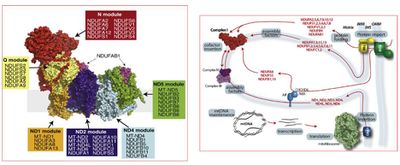
Figure 1: (left) modular composition of mt CI; (right) Assembly pathway of mt CI4.
References
- Gnaiger E, Aasander Frostner E, Abdul Karim N, Abumrad NA, Acuna-Castroviejo D, Adiele RC, et al (2019) Mitochondrial respiratory states and rates. MitoFit Preprint Arch doi:10.26124/mitofit:190001.v4.
- [www.um.edu.mt/biobank]
- [www.platform.rd-connect.eu]
- Formosa LE, Dibley MG, Stroud DA, Ryan MT (2018) Building a complex complex: assembly of mitochondrial respiratory chain complex I. Semin Cell Dev Biol 76:154-62.