Musicco 2023 Int J Mol Sci
| Musicco C, Signorile A, Pesce V, Loguercio Polosa P, Cormio A (2023) Mitochondria deregulations in cancer offer several potential targets of therapeutic interventions. Int J Mol Sci 24:10420. https://doi.org/10.3390/ijms241310420 |
Musicco C, Signorile A, Pesce V, Loguercio Polosa P, Cormio A (2023) Int J Mol Sci
Abstract: Mitochondria play a key role in cancer and their involvement is not limited to the production of ATP only. Mitochondria also produce reactive oxygen species and building blocks to sustain rapid cell proliferation; thus, the deregulation of mitochondrial function is associated with cancer disease development and progression. In cancer cells, a metabolic reprogramming takes place through a different modulation of the mitochondrial metabolic pathways, including oxidative phosphorylation, fatty acid oxidation, the Krebs cycle, glutamine and heme metabolism. Alterations of mitochondrial homeostasis, in particular, of mitochondrial biogenesis, mitophagy, dynamics, redox balance, and protein homeostasis, were also observed in cancer cells. The use of drugs acting on mitochondrial destabilization may represent a promising therapeutic approach in tumors in which mitochondrial respiration is the predominant energy source. In this review, we summarize the main mitochondrial features and metabolic pathways altered in cancer cells, moreover, we present the best known drugs that, by acting on mitochondrial homeostasis and metabolic pathways, may induce mitochondrial alterations and cancer cell death. In addition, new strategies that induce mitochondrial damage, such as photodynamic, photothermal and chemodynamic therapies, and the development of nanoformulations that specifically target drugs in mitochondria are also described. Thus, mitochondria-targeted drugs may open new frontiers to a tailored and personalized cancer therapy.
• Bioblast editor: Gnaiger E
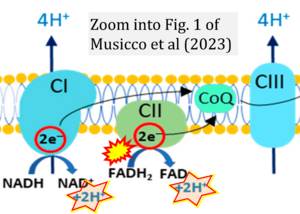
Correction: FADH2 and Complex II
- FADH2 is shown as the substrate feeding electrons into Complex II (CII). This is wrong and requires correction - for details see Gnaiger (2024).
- Gnaiger E (2024) Complex II ambiguities ― FADH2 in the electron transfer system. J Biol Chem 300:105470. https://doi.org/10.1016/j.jbc.2023.105470 - »Bioblast link«
Hydrogen ion ambiguities in the electron transfer system
Communicated by Gnaiger E (2023-10-08) last update 2023-11-10
- Electron (e-) transfer linked to hydrogen ion (hydron; H+) transfer is a fundamental concept in the field of bioenergetics, critical for understanding redox-coupled energy transformations.
- However, the current literature contains inconsistencies regarding H+ formation on the negative side of bioenergetic membranes, such as the matrix side of the mitochondrial inner membrane, when NADH is oxidized during oxidative phosphorylation (OXPHOS). Ambiguities arise when examining the oxidation of NADH by respiratory Complex I or succinate by Complex II.
- Oxidation of NADH or succinate involves a two-electron transfer of 2{H++e-} to FMN or FAD, respectively. Figures indicating a single electron e- transferred from NADH or succinate lack accuracy.
- The oxidized NAD+ is distinguished from NAD indicating nicotinamide adenine dinucleotide independent of oxidation state.
- NADH + H+ → NAD+ +2{H++e-} is the oxidation half-reaction in this H+-linked electron transfer represented as 2{H++e-} (Gnaiger 2023). Putative H+ formation shown as NADH → NAD+ + H+ conflicts with chemiosmotic coupling stoichiometries between H+ translocation across the coupling membrane and electron transfer to oxygen. Ensuring clarity in this complex field is imperative to tackle the apparent ambiguity crisis and prevent confusion, particularly in light of the increasing number of interdisciplinary publications on bioenergetics concerning diagnostic and clinical applications of OXPHOS analysis.
Labels: MiParea: Pharmacology;toxicology Pathology: Cancer