Difference between revisions of "SUIT-007"
From Bioblast
Beno Marija (talk | contribs) |
Beno Marija (talk | contribs) |
||
| (One intermediate revision by the same user not shown) | |||
| Line 1: | Line 1: | ||
{{MitoPedia | {{MitoPedia | ||
|abbr=[[Glutamate anaplerotic pathway control state|Glutamate anaplerosis]] | |abbr=[[Glutamate-anaplerotic pathway control state|Glutamate anaplerosis]] | ||
|description=[[File:1G;2D;3M;4U-.png|300px]] | |description=[[File:1G;2D;3M;4U-.png|300px]] | ||
|info='''A:''' '''[[Glutamate anaplerotic pathway control state|Glutamate anaplerotic pathway]]''' | |info='''A:''' '''[[Glutamate-anaplerotic pathway control state|Glutamate anaplerotic pathway]]''' | ||
}} | }} | ||
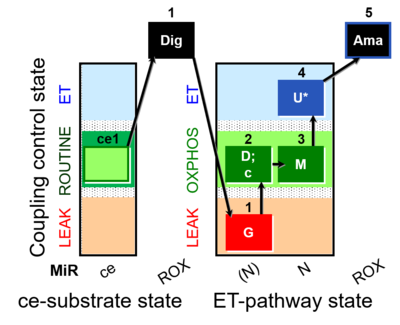
::: '''[[SUIT protocol pattern]]:''' 1G;2D;2c;3M;4U;5Ama | ::: '''[[SUIT protocol pattern]]:''' 1G;2D;2c;3M;4U;5Ama | ||
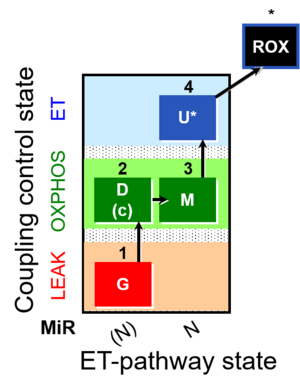
The SUIT-007 is focused on the [[Glutamate anaplerotic pathway control state|glutamate anaplerotic pathway]]. If [[Glutamate dehydrogenase|glutamate dehydrogenase]] is present in the sample, when glutamate alone is added, it will be converted to α-ketoglutarate in an [[Anaplerosis|anaplerotic reaction]], supporting respiration. After malate is added, it is possible to analyse the [[NADH Electron transfer-pathway state]] with glutamate and malate as substrates in the [[OXPHOS]] and [[ | The SUIT-007 is focused on the [[Glutamate-anaplerotic pathway control state|glutamate anaplerotic pathway]]. If [[Glutamate dehydrogenase|glutamate dehydrogenase]] is present in the sample, when glutamate alone is added, it will be converted to α-ketoglutarate in an [[Anaplerosis|anaplerotic reaction]], supporting respiration. After malate is added, it is possible to analyse the [[NADH Electron transfer-pathway state]] with glutamate and malate as substrates in the [[OXPHOS]] and [[Electron transfer pathway]] [[Coupling control state|coupling control states]]. | ||
| Line 20: | Line 20: | ||
== Strengths and limitations == | == Strengths and limitations == | ||
:::* Protocol focused on analysis of glutamate anaplerotic pathway/glutaminolysis. | :::* Protocol focused on analysis of glutamate anaplerotic pathway/glutaminolysis. | ||
:::+ The use of glutamate alone allows to analyse the glutamate anaplerotic pathway control state. | :::+ The use of glutamate alone allows to analyse the glutamate-anaplerotic pathway control state. | ||
:::+ Short duration of the experiment. | :::+ Short duration of the experiment. | ||
:::+ Glutamate is easier to prepare compared to pyruvate. | :::+ Glutamate is easier to prepare compared to pyruvate. | ||
Latest revision as of 11:30, 3 June 2020
Description
Abbreviation: Glutamate anaplerosis
Reference: A: Glutamate anaplerotic pathway
- SUIT protocol pattern: 1G;2D;2c;3M;4U;5Ama
The SUIT-007 is focused on the glutamate anaplerotic pathway. If glutamate dehydrogenase is present in the sample, when glutamate alone is added, it will be converted to α-ketoglutarate in an anaplerotic reaction, supporting respiration. After malate is added, it is possible to analyse the NADH Electron transfer-pathway state with glutamate and malate as substrates in the OXPHOS and Electron transfer pathway coupling control states.
Communicated by Cardoso LH, Komlodi T, Doerrier C and Gnaiger E (last update 2019-02-25)
Specific SUIT protocols
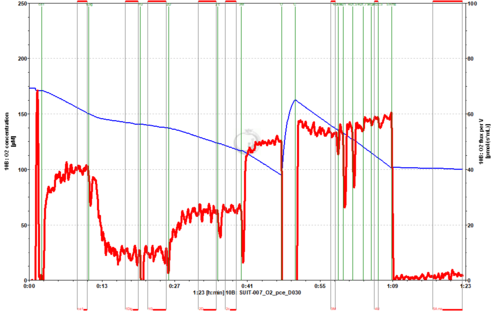
- SUIT-007 O2 ce-pce D030 for permeabilized cells
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1G | GL | (N) | CI | 1G
|
| 2D | GP | (N) | CI | 1G;2D
|
| 2c | GcP | (N) | CI | 1G;2D;2c
|
| 3M | GMP | N | CI | 1G;2D;2c;3M
|
| 4U | GME | N | CI | 1G;2D;2c;3M;4U
|
| 5Ama | ROX | 1G;2D;2c;3M;4U;5Ama
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- Protocol focused on analysis of glutamate anaplerotic pathway/glutaminolysis.
- + The use of glutamate alone allows to analyse the glutamate-anaplerotic pathway control state.
- + Short duration of the experiment.
- + Glutamate is easier to prepare compared to pyruvate.
- + CIV activity can be measured after this protocol.
- - Analysis of N-OXPHOS capacity (instead of NS-OXPHOS capacity) does not provide the best estimate of maximum mitochondrial respiratory capacity.
- - This protocol does not address questions regarding the convergence of pathways at the Q-junction.
- - Addition of cytochrome c after glutamate alone (1G;2D;2c) is not as appropriated to evaluate integrity of the mitochondrial external membrane as it would be with GM. However, with addition of cytochrome c only after malate (e.g. 1G;2D;3M;3c), it would not be possible to compare GP and GMP if there are damages to the mitochondrial external membrane, therefore it is suggested to add cytochrome c after ADP (1G;2D;2c).
- - Careful washing is required after the experiment to avoid carry-over of inhibitors and uncoupler.
Compare SUIT protocols
- SUIT-006: Coupling-control protocol, can be performed with GM.
References
MitoPedia concepts: MiP concept, SUIT protocol, Recommended
MitoPedia methods:
Respirometry