Difference between revisions of "SUIT-022"
From Bioblast
| Line 6: | Line 6: | ||
::: '''[[SUIT protocol pattern]]:''' linear ce1;ce2KCN;ce3SHAM | ::: '''[[SUIT protocol pattern]]:''' linear ce1;ce2KCN;ce3SHAM | ||
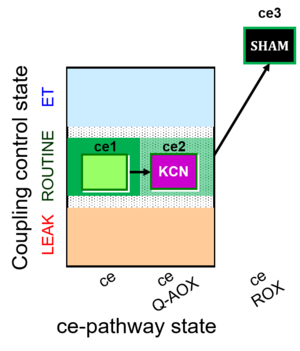
The SUIT-022 is designed to provide an estimation of the mitochondrial respiration in cells due to the [[alternative oxidase]] pathway, which is [[cyanide| cyanide]]-and [[Antimycin_A| antimycin A]]-resistant. This protocol must be complemented with the study in parallel of the mitochondrial respiration due to the cytochrome ''c'' pathway (see [[SUIT-023]]). | The SUIT-022 is designed to provide an estimation of the mitochondrial respiration in cells due to the [[alternative oxidase]] pathway, which is [[cyanide| cyanide]]-and [[Antimycin_A| antimycin A]]-resistant. This protocol must be complemented with the study in parallel of the mitochondrial respiration due to the cytochrome ''c'' oxidase pathway (see [[SUIT-023]]). | ||
| Line 19: | Line 19: | ||
* [[SUIT-022 O2 ce D051]] for intact microalgal cells (ce) | * [[SUIT-022 O2 ce D051]] for intact microalgal cells (ce) | ||
{{Template:SUIT-022}} | {{Template:SUIT-022 O2 ce D00}} | ||
== Strengths and limitations == | == Strengths and limitations == | ||
Revision as of 11:34, 9 April 2019
Description
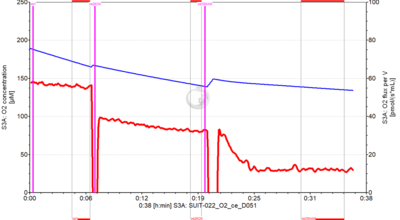
Reference: A: Determination of the respiration due to the alternative oxidase pathway ![]() »Versions
»Versions
O2k-Application: O2
- SUIT-category: ce
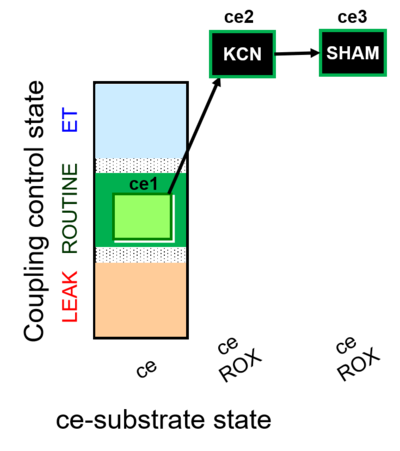
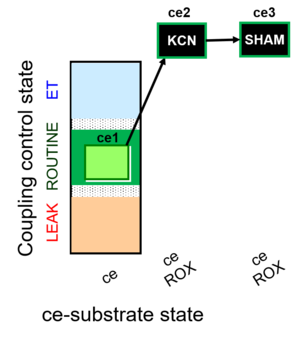
- SUIT protocol pattern: linear ce1;ce2KCN;ce3SHAM
The SUIT-022 is designed to provide an estimation of the mitochondrial respiration in cells due to the alternative oxidase pathway, which is cyanide-and antimycin A-resistant. This protocol must be complemented with the study in parallel of the mitochondrial respiration due to the cytochrome c oxidase pathway (see SUIT-023).
Communicated by Huete-Ortega M and Gnaiger E (last update 2019-04-09)
Specific SUIT protocols
SUIT-022 O2 ce D051
- SUIT-022 O2 ce D051 for intact microalgal cells (ce)
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| ce1 | ROUTINE | ce1
| ||
| ce2KCN | ROX or ROUTINE | ce1;ce2KCN
| ||
| ce3SHAM | ROX | ce1;ce2KCN;ce3SHAM
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- SUIT-022 provides an estimation of the cyanide-and antimycin A-resistant mitochondrial respiration due to the activity of the alternative oxidase on cells on ROUTINE. To complement the study with the estimation of the mitochondrial respiration due to the cytochrome c pathway, it must be carried out in parallel to SUIT-023.
- + This protocol can be used with many type of cells, although optimal concentrations of inhibitors have to be optimised.
- + Short duration of experiment
- - Estimation of the linear coupling control is not done
- - The use of cyanide requires a careful and in deep cleaning of the chambers after use and careful handling of the chemical to guarantee health safety.
Compare SUIT protocols
- SUIT-023 O2 ce D053 for microalgal intact cells
References
MitoPedia concepts: MiP concept, SUIT protocol, Recommended
MitoPedia methods:
Respirometry