Simon 2022 Function (Oxf)
| Simon L, Molina PE (2022) Cellular bioenergetics: experimental evidence for alcohol-induced adaptations. Function (Oxf) 3:zqac039. https://doi.org/10.1093/function/zqac039 |
Simon L, Molina PE (2022) Function (Oxf)
Abstract: At-risk alcohol use is associated with multisystemic effects and end-organ injury, and significantly contributes to global health burden. Several alcohol-mediated mechanisms have been identified, with bioenergetic maladaptation gaining credence as an underlying pathophysiological mechanism contributing to cellular injury. This evidence-based review focuses on the current knowledge of alcohol-induced bioenergetic adaptations in metabolically active tissues: liver, cardiac and skeletal muscle, pancreas, and brain. Alcohol metabolism itself significantly interferes with bioenergetic pathways in tissues, particularly the liver. Alcohol decreases states of respiration in the electron transport chain, and activity and expression of respiratory complexes, with a net effect to decrease ATP content. In addition, alcohol dysregulates major metabolic pathways, including glycolysis, the tricarboxylic acid cycle, and fatty acid oxidation. These bioenergetic alterations are influenced by alcohol-mediated changes in mitochondrial morphology, biogenesis, and dynamics. The review highlights similarities and differences in bioenergetic adaptations according to tissue type, pattern of (acute vs. chronic) alcohol use, and energy substrate availability. The compromised bioenergetics synergizes with other critical pathophysiological mechanisms, including increased oxidative stress and accelerates cellular dysfunction, promoting senescence, programmed cell death, and end-organ injury.
• Bioblast editor: Gnaiger E
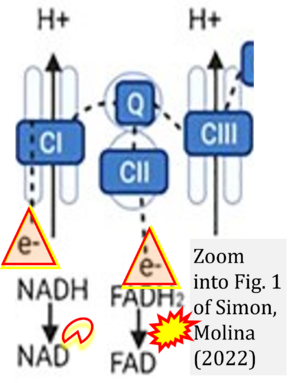
Correction: FADH2 and Complex II
- FADH2 is shown as the substrate feeding electrons into Complex II (CII). This is wrong and requires correction - for details see Gnaiger (2024).
- Gnaiger E (2024) Complex II ambiguities ― FADH2 in the electron transfer system. J Biol Chem 300:105470. https://doi.org/10.1016/j.jbc.2023.105470 - »Bioblast link«
Hydrogen ion ambiguities in the electron transfer system
Communicated by Gnaiger E (2023-10-08) last update 2023-11-10
- Electron (e-) transfer linked to hydrogen ion (hydron; H+) transfer is a fundamental concept in the field of bioenergetics, critical for understanding redox-coupled energy transformations.
- However, the current literature contains inconsistencies regarding H+ formation on the negative side of bioenergetic membranes, such as the matrix side of the mitochondrial inner membrane, when NADH is oxidized during oxidative phosphorylation (OXPHOS). Ambiguities arise when examining the oxidation of NADH by respiratory Complex I or succinate by Complex II.
- Oxidation of NADH or succinate involves a two-electron transfer of 2{H++e-} to FMN or FAD, respectively. Figures indicating a single electron e- transferred from NADH or succinate lack accuracy.
- The oxidized NAD+ is distinguished from NAD indicating nicotinamide adenine dinucleotide independent of oxidation state.
- NADH + H+ → NAD+ +2{H++e-} is the oxidation half-reaction in this H+-linked electron transfer represented as 2{H++e-} (Gnaiger 2023). Putative H+ formation shown as NADH → NAD+ + H+ conflicts with chemiosmotic coupling stoichiometries between H+ translocation across the coupling membrane and electron transfer to oxygen. Ensuring clarity in this complex field is imperative to tackle the apparent ambiguity crisis and prevent confusion, particularly in light of the increasing number of interdisciplinary publications on bioenergetics concerning diagnostic and clinical applications of OXPHOS analysis.
Labels: MiParea: Pharmacology;toxicology
Enzyme: Complex II;succinate dehydrogenase
HRR: Oxygraph-2k
EtOH