Alston 2017 J Pathol
| Alston CL, Rocha MC, Lax NZ, Turnbull DM, Taylor RW (2017) The genetics and pathology of mitochondrial disease. J Pathol 241:236-50. https://doi.org/10.1002/path.4809 |
Alston CL, Rocha MC, Lax NZ, Turnbull DM, Taylor RW (2017) J Pathol
Abstract: Mitochondria are double-membrane-bound organelles that are present in all nucleated eukaryotic cells and are responsible for the production of cellular energy in the form of ATP. Mitochondrial function is under dual genetic control - the 16.6-kb mitochondrial genome, with only 37 genes, and the nuclear genome, which encodes the remaining ∼1300 proteins of the mitoproteome. Mitochondrial dysfunction can arise because of defects in either mitochondrial DNA or nuclear mitochondrial genes, and can present in childhood or adulthood in association with vast clinical heterogeneity, with symptoms affecting a single organ or tissue, or multisystem involvement. There is no cure for mitochondrial disease for the vast majority of mitochondrial disease patients, and a genetic diagnosis is therefore crucial for genetic counselling and recurrence risk calculation, and can impact on the clinical management of affected patients. Next-generation sequencing strategies are proving pivotal in the discovery of new disease genes and the diagnosis of clinically affected patients; mutations in >250 genes have now been shown to cause mitochondrial disease, and the biochemical, histochemical, immunocytochemical and neuropathological characterization of these patients has led to improved diagnostic testing strategies and novel diagnostic techniques. This review focuses on the current genetic landscape associated with mitochondrial disease, before focusing on advances in studying associated mitochondrial pathology in two, clinically relevant organs - skeletal muscle and brain. © 2016 The Authors. The Journal of Pathology published by John Wiley & Sons Ltd on behalf of Pathological Society of Great Britain and Ireland.
• Bioblast editor: Gnaiger E
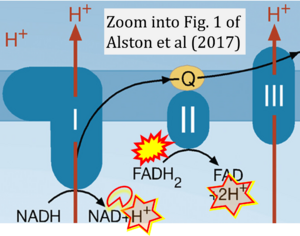
Correction: FADH2 and Complex II
- FADH2 is shown as the substrate feeding electrons into Complex II (CII). This is wrong and requires correction - for details see Gnaiger (2024).
- Gnaiger E (2024) Complex II ambiguities ― FADH2 in the electron transfer system. J Biol Chem 300:105470. https://doi.org/10.1016/j.jbc.2023.105470 - »Bioblast link«
Hydrogen ion ambiguities in the electron transfer system
Communicated by Gnaiger E (2023-10-08) last update 2023-11-10
- Electron (e-) transfer linked to hydrogen ion (hydron; H+) transfer is a fundamental concept in the field of bioenergetics, critical for understanding redox-coupled energy transformations.
- However, the current literature contains inconsistencies regarding H+ formation on the negative side of bioenergetic membranes, such as the matrix side of the mitochondrial inner membrane, when NADH is oxidized during oxidative phosphorylation (OXPHOS). Ambiguities arise when examining the oxidation of NADH by respiratory Complex I or succinate by Complex II.
- Oxidation of NADH or succinate involves a two-electron transfer of 2{H++e-} to FMN or FAD, respectively. Figures indicating a single electron e- transferred from NADH or succinate lack accuracy.
- The oxidized NAD+ is distinguished from NAD indicating nicotinamide adenine dinucleotide independent of oxidation state.
- NADH + H+ → NAD+ +2{H++e-} is the oxidation half-reaction in this H+-linked electron transfer represented as 2{H++e-} (Gnaiger 2023). Putative H+ formation shown as NADH → NAD+ + H+ conflicts with chemiosmotic coupling stoichiometries between H+ translocation across the coupling membrane and electron transfer to oxygen. Ensuring clarity in this complex field is imperative to tackle the apparent ambiguity crisis and prevent confusion, particularly in light of the increasing number of interdisciplinary publications on bioenergetics concerning diagnostic and clinical applications of OXPHOS analysis.
Labels:
Enzyme: Complex II;succinate dehydrogenase