Talk:Oligomycin
From Bioblast
MiPNet discussion forum: washing out Omy (2013-12-11)
Stine Lundby
- Retention of inhibitors (Oligomycin, I assume) in the chambers. Here is the situation: all four chambers work close to perfect! Permeabilized human muscle fibers respire superb in the morning (biopsy taken in the morning). Permeabalized human muscle fibers respire 3-fold lower in the afternoon (biopsy taken in the afternoon) – FCCP response as in the morning.
- Included inhibitors in the SUIT: Rot, AmA and OMY.
- Wash between runs: 3*5 min H2O, 20 min yeast, 3*5 min 70% EtOH, 30 min 100% EtOH,3*5 min water.
- Any clue of what we can do to get rid of the inhibitors when running experiments in the morning and in the afternoon?
Douglas L Crawford
- The simplest and most effect method for removing oligomycin is to wash chamber, then put cell/mito into the chamber for a few minutes. That is, use previous cell/mito preparation (I keep excessive mitts in the freezer) to absorb the trace amount of Oligomycin, then wash the chamber again.
- 2nd: use less oligomycin.
Ib Therkelsen Arent
- Concering the Oligomycin situation. I assume that you also rinse the stoppers properly. As to the rinsing process of the chambers, I do not think that the time involved means anything, but rather the degree of dilution, i.e. wash MANY times with water, alcohol, and water again (can be done within a few minutes) . I don't see that the yeast should help in any way.
- Biofilm may build up on the glass walls in time, and this has earlier on been a nuisance in our lab. Between runs, a cleaning of the chamber walls with cotton sticks (ear pickers, or whatever they are called) might do the trick.
- Working with a variety of poisons here: Oligomycin, Antimycin-A, Rotenone, NaN3, KCN (on isolated mitochondria, mostly, for the time being). Never had a problem with any of these. However, we DO NOT use FCCP any longer. Results were were often capricious and unreliable.
- Are the biopsies prepared for measurements by the same person morning and afternoon? Is the Saponin (relatively) freshly prepared?
- Best of luck from Ib Th. Arent, Copenhagen
Magda Labieniec-Watala
- About 2 years ago I had similar problems with mito inhibitors, but in those days I was carrying out the experiments for a long time during the day (about 10-12 hours), started at 9 a.m. and finishing at 9-10 p.m. Then, the appropriate chamber washing procedure helped a lot. It means: 3 * H2O, 3 * 70% ethanol, 1 * 70% ethanol for 10-15 min, 3 * H2O, 1 * 96% ethanol for 15 min; 3 * H2O. Nevertheless, your 'wash runs' seem enough. In my opinion it is very likely that the difference in the sample taking - (morning/evening) - has the impact on mitochondrial bioenergetics. Probably, the differences in the organism metabolism are reflected in your measurements rather than the use of inhibitors. Please, start your experiments in the evening to verify this hypothesis :), if it is possible.
Dominik Pesta
- I am aware of this problem and we had similar issues. When I wash - and your washing procedure is more extensive than mine - I do not have these problems and respiration is back. The critical wash for me is with EtOH absolut, though. And I simply stopped using Rotenone (with only 1 oxy washing would slow me down too much).
- Stuff to do: (1) as addressed above, make sure to insert the stoppers tightly into the EtOH during washing, chemicals might get stuck in the rubber rings.
- (2) remove chambers and soak and clean thoroughly in H2O2 (with those cotton sticks as Ib mentioned). A biofilm on the chamber might contribute to chemical-retention....
- (3) As a last resort (if it's really the chambers): get an extra pair of glass chambers and change them after the first run (and soak them overnight) while inserting the "fresh" ones. Might give you some issues with BG but it might be worth a try... Good luck!!
Malou Friederich Persson
- Previously I also had trouble with using oligomycin in one chamber staying also in the second run, lowering the RCR substantially compared to if I ran in the chamber that hadn´t contained oligomycin in during the first experiment. At first I just cleaned with ethanol for a LONG time (3x5 minutes + 3x20 minutes!) but then I got a tip of adding a lot of protein to the chamber to help clean the oligomycin.
- So now, I add either left over mitochondria (about 200 microliter of a fairly concentrated stock (the chamber window should show a solution pretty clouded with protein). Sometimes, if I have a shortage of mitochondria (I isolate from kidney tissue) I save some crude tissue homogenate (the pellet containing nuclei and unbroken tissue from the first centrifugation step during the isolation process) and use that. It does not matter what kind of protein you use, as long as it is a lot of it! The more the better. Just add protein, let sit for 2-3 minutes and wash 5-6 times with water and then buffer. Also, put the lid on all the way down when the protein is in, otherwise the oligomycin may still stick to the lid.
- With this procedure I have no problem at all getting rid of oligomycin in about 5 minutes and I save a lot of time in between runs.
- This should work for all ethanol-based inhibitors/substances I guess.
Antonio Galina
- I usually wash the chamber with a frozen liver mitochondrial preparation. I put about 1 mg/mL of rat liver mitochondria and incubate for 5 minutes and then 5 times absolute ETOH, and more 5 times with distilled water. Using this procedure I did not have any contamination problems with inhibitors or uncouplers.
Marcus Oliveira
- This is possibbly due to remnants of OXPHOS inhibitors, rotenone, antimycin, oligomycin, within the chambers, despite extensive water and ethanol washings.
- Try to wash the chambers, after finishing a run, with rat liver homogenate previously prepared. This usually improves the removal of inhibitors and, in our hands, eliminated these problems.
Paula M. Keeney
- To wash out the oligomycin try adding 20 min in 5% Na2SO4 (sodium sulfite) the wash between runs. We would then wash the chamber with 5 x H20, 5 x 70% 5 x 100% EtOH and 5 x water.
MiPNet discussion forum: problem with Omy (2013-10-22)
Steffi Wohlgemuth
- The problem concerns the unresponsiveness of our cells, isolated mitos, and permeabilized muscle cells to oligomycin. When we titrate oligomycin to our cells or to our permeabilized muscle fibers (and the same happened with isolated mitos in the past in another lab) to induce state 4 or leak respiration, the flux only decreases by a little bit, sometimes returns to the flux before oligomycin-addition after a short while. -- I have abandoned the O2k for a few months and want to pick up those experiments again, but my students are reluctant to run the oligomycin-induced state because it has almost never worked and prolongs the protocol with no added information. -- I am sure I will be asked what concentration of Oligo we added: The recommended concentration of 2 ug/mL of a 4mg/1mL EtOH stock. We even started making a little bit more dilute Oligo stock in order to titrate smaller amounts up to the final concentration to see if less oligo would do the trick, if we added to much, and if our cells were more sensitive and less oligo would be required. -- No satisfying results.
- So my overall question: what is the deal with Oligo? I have heard from others as well that oligo is not quite working as it is supposed to. What can we do differently? Are we doing anything wrong? Is there a better oligo besides the one recommended from Sigma.... Sometimes the batches are different or the manufacturer changes procedures.....
- Any help or suggestion at this point would be awesome and very, very welcomed. [[email protected]]
Felix Mark
- We never had fundamental issues with oligomycin, but in our assays with temperate and polar fish and cephalopod permeabilized fibres (heart muscle), it sometimes takes ages for a result to become visible and stable. So it can depend on assay temperature and time, as well. At low temperatures, eg. 0-5°C, we sometimes wait up to 45min.
- Have you tried atractyloside? It's not quite the same response as it blocks the ANT rather than ATPase, but should give similar results in terms of flux reduction and membrane potential. And, at least in our preps, it's much faster. We sometimes use it to double-check.
- I assume you have also tried preparing a new stock solution from a fresh new batch of oligomycin, that's usually our last resort…. [[email protected]]
Marten Wikstrom
- I would recommend the following:
- First test whether your oligomycin will block the ADP (+ Pi)-induced increase in respiratory rate (State 4-->State 3) with isolated mitochondria respiring on succinate. That will test whether your oligo batch is working/whether you use a sufficient concentration of oligo.
- If the test is positive (oligomycin blocks the State 4->State 3 transition), then to understand the unresponsiveness to oligo in other experiments would require detailed knowledge of the conditions in those expts.
- If the test is negative, throw your oligo batch away and get a new one. [[email protected]]
Senyilmaz, Deniz
- What is the brand of your oligomycin? Initially I had the same problem exactly, but then I switched to Sigma, and oligomycin started to work! For some reason, the other brand failed to induce state 4 respiration even though it was able to activate other downstream processes in the cultured cells. [[email protected]]
Lars Eide
- Must be your oligomycin batch: we actually experience the opposite problem: oligomycin is hard to get rid of, and extensive wash (of an 7-years old oxygraph) is required to reset the system for next run. [[email protected]]
Christopher Hedges
- I have worked with oligomycin in both the Stepto lab in Melbourne and the Hickey lab in Auckland. In Melbourne we have had extensive trouble with uncoupling following oligo (our uncoupled respiration was frequently less than oxphos in permeabilised muscle preps). I haven't personally experienced what you describe with respiration returning to prior levels but certainly had trouble with oligo - ultimately our solution was to stop using oligo and use atractyloside or carboxyatractyloside instead. This may or may not suit what you need to do. An important consideration is that ANT conducts some protons so contributes to leak respiration. We have found that adding atractyloside after oligomycin causes another small decrease in oxygen consumption, even though they theoretically both induce the same state (high substrate, low ADP turnover). I have not used permeabilised cells and my limited work with isolated mito's we had already given up on oligo so this was all found with permeabilised skeletal muscle fibers. [[email protected]]
Michael Duchen
- Oligomycin can be terribly 'sticky' - I have had the experience years ago of using imaging dishes (for which the coverslip on which cells were grown formed the base) which had been used for oligomycin previously and the cells behaved as though they had already seen oligo. As it is effectively irreversible is it conceivable that you don't see responses because the cells have effectively already been exposed? It is necessary to wash components very carefully with ethanol.
- To test whether oligo has been effective or not, the easiest thing to do is to measure the potential and see how the potential responds to inhibition of respiration using rotenone or cyanide. Under control conditions, depolarisation is rather slow in most cells, but is much faster and more complete after exposure to oligomycin as the ATPase can't run in reverse to maintain potential.
- I can't think of anything else other than try a new batch of oligomycin! [[email protected]]
Alessandro Pontoglio
- My name is Alessandro Pontoglio, it's three years since I worked and graduated as "Dottore in ricerca" (Doctorate, Ph.D) in the team of Prof. Scatena Roberto, at Policlinico A. Gemelli in Rome, and what I managed was the O2K techincs and technologies applied to hepatocarcinoma cells (HepG2) cultures permeabilized and non permeabilized. Sometimes promielocitic leukemia cells (HL60) cultures.
- As I've read in your message : your team is experiencing some troubles with oligomycin flux determination.
- I've read you're using muscle cells permeabilized and no permeabilized, and even mitochondria from the same kind of cells. Am I right?
- Would you please answer to these questions? It would be worthy in order to try to understand what can be improved or changed. Thank you.
- NO PERMEABILIZED CELLS :
- # What are the concentration in O2K chamber, the origin, the age, the number of changes of these cells (if cultured for a long time)?
- # What's the basal flux respiration? Do you think it is already low at the beginning of the determination or it is normal if compared to what is written in the literature?
- # Is the experimental pathway of your experiments : basal-oligomicym-FCCP-Rotenone-Antimycin? Or else? If you think you can disclose them, would you show me the values of the corresponding respiratory fluxes?
- NO PERMEABILIZED CELLS :
- PERMEABILIZED CELLS :
- # What's the cell concentration in the O2K chambers?
- # The experimental pathway your team performs is : basal-(malate+glutammate)-digitonin or saponin-ADP-Succinate-ADP-ADP-oligomycin-FCCP-Rotenone-Malonate-Antimycin or another?
- # If any, have you ever seen well shaped spikes when ADP is added, e.g. with succinate?
- PERMEABILIZED CELLS :
- GENERAL : Supposing the concentration of oligomycin your team started with was the right one :
- # Have you already performed the same experiments on other cells suh as HepG2? They were very responsive to reactants used. But if I tell you about HL60...they had a very low basal flux (their normal respiration) and the responsiveness to oligomycin and FCCP was very restrained.
- # Is the ATP dependent respiratory activity of the tested cells significative?
- # No ATPase inhibitor substances or uncoupling agents are in the suspension media used or bacteria in cultures?
- # What about cells and mitochondria integrity during and after isolation and treatment of these two subjects?
- [[email protected]]
- GENERAL : Supposing the concentration of oligomycin your team started with was the right one :
Ib Therkelsen Arent
- In our lab, a stock solution of 200 µM in 96% EtOH is kept in the freezer for months on end.
- To each Oxygraph chamber (2 ml) is added 5 µl during experiments, resulting in a final conc of 0,5 µM.
- With Isolated Mitos we never had a problem.
- With fibers I don't recall if we have ever used it.
- The Producers name I am not familiar with, but I might ask someone. [[email protected]]
Adam Chicco
- You might have ruled this out already, but we see the same in muscle/heart mitochondria and fibers when adding oligomycin in the presence of fatty acid substrate (e.g., palmitoylcarnitine), probably due to uncoupling effects of the fatty acids. So, if fatty acids are a standard part of your assays or prep, I'd test the same oligomycin with glutamate or pyruvate+malate alone to see if you get more inhibition. [[email protected]]
Peter Schoenfeld
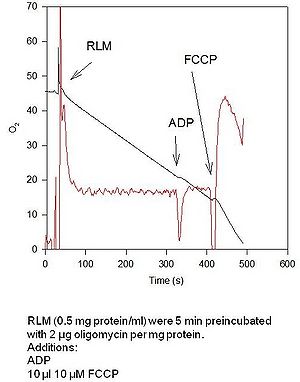
- When mitochondria were shortly preincubated with oligomycin, there is an inhibition (see Figure). For your information, an experiment is added.
- In this experiment a 1 mM ethanolic oligomycin solution was applied. The oxygen uptake was measured with an oxygraph from the first generation. [[email protected]]
Frédéric Bouillaud
- First I do apologize for the trivial aspect of my answer.
- It happened to us several times and in our hands it turned out that the explanation was that there was a contamination with low amount of oligomycin.
- A very small amount of oligomycin left in the chamber (stopper) is suffcient to develop a complete inhibition because exposure to it took place for minutes (setting of exp and equilibration). Of course once inhibited they do not respond to it anymore, increasing oligo input in fact aggravates the problem.
- With cells you can't probe for ADP response with mitochondria you could.
- If you don't wait for equilibration and close the chamber immediately you'll observe the decline in respiratory rate.
- One should realize that the respiratory rate is low in terms of pmol/(s.ml) but in fact with cells preps you often don't know what is the "correct value". [[email protected]]
ET capacity after inhibition by Omy (2013-02-21)
- * We are having problems recovering full activity with FCCP after oligomycin, and although diluting Omy to low but effective concentration help, it does not solve the problem. Thus, we are considering buying the more expensive forms of Omy. (2013-02-21; Doug Crawford, University of Miami, US)
- * We are facing the same problems during recent years, and are worried about what is going wrong. Whereas pathologically stressed mitos with a latent injury do respond to the additional Omy induced stress by loosing ET capacity, the controls gave us recently the same problems, which we consider as artefacts for unknown reasons. Quality control of formerly functional chemicals becomes an increasingly worrying issue. (Erich Gnaiger, Medical University Innsbruck, AT)
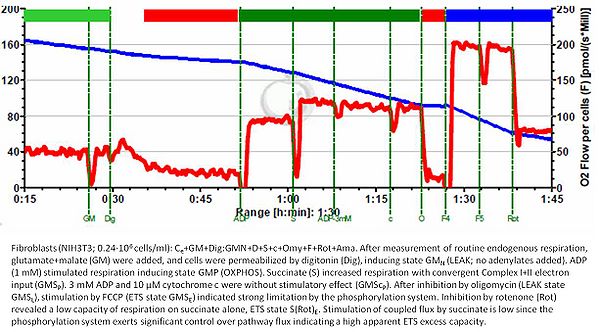
- * Here is an example where ET capacity was not impaired after Omy treatment. In these NIH3T3 fibroblasts ET capacity clearly exceeded OXPHOS capacity and this was similar to controls measured in the absence of Omy (not shown). The example shown here is from Gnaiger 2020 BEC MitoPathways.
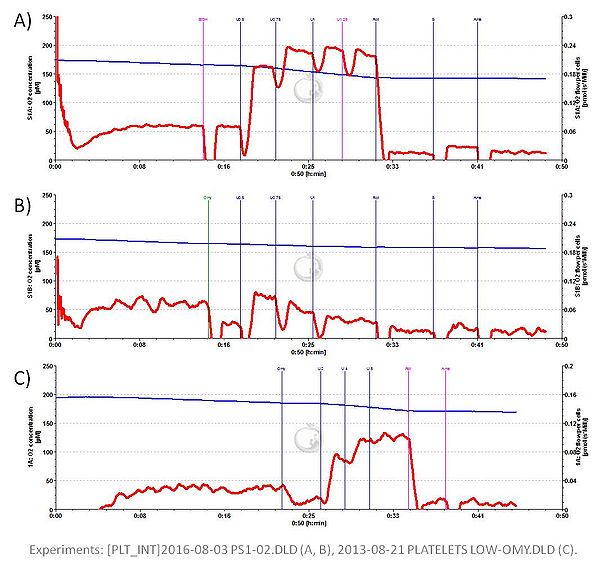
- * A possible way of overcoming/circumventing the problem of Omy inhibiting subsequently measured ET capacity has been suggested by Garcia-Souza Luiz F and colleagues. Measuring respiration in intact human platelets a merely 1.2-fold increase of oxygen flux from ROUTINE to apparent ET capacity was detected after Omy addition, whereas a 2.7-fold increase was observed after adding only the solvent ethanol. Now by simply reducing Omy concentration to one fifth (from 2.5 µM to 0.5 µM) the 2.7-fold increase was fully preserved while leaving the reduction from ROUTINE to LEAK respiration unaltered.
- Respiration in intact isolated human platelets; experimental buffer: RPMI + 2 mM glutamine (Figs A and B) and M199 (Fig. C); 37 C. Oligomycin (Omy) was added at 2.5 µM (Fig A) and 0.5 µM (Fig. C), while only ethanol was added to cells in Fig. B. CCCP (U) was added in 0.2-0.5 µM steps; rotenone (Rot) at 1 µg/ml, antimycin A (Ama) at 2.5µM final concentration.
Lack of inhibition with Omy in yeast cells (2014-08-07)
- (Contributed by Gennaro Agrimi)
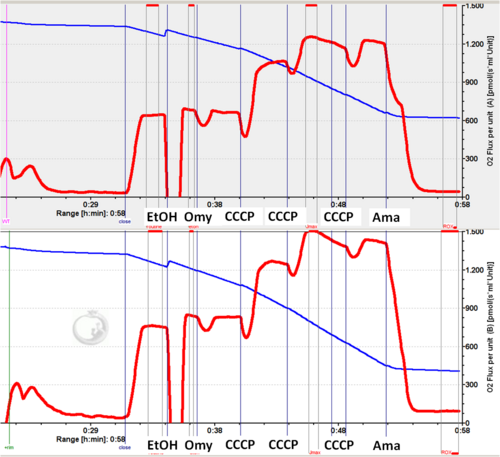
- Exponentially growing cultures of S. cerevisiae strain BY4742 in YP supplemented with 3% glycerol were harvested at an OD600 of 0.7 to 1, centrifuged at 3,000 x g for 5 min at 4°C, and resuspended in YP at a density of approximately 5 OD600 units. 50 μl of this suspension corresponding to about 5 × 106 cells/ml were added to each chamber containing 2 ml of YP. The chambers were closed and Routine respiration was recorded.
- Oligomycin was purchased from Santa Cruz Biotechnology (SC-201551 - Lot. # A3114). 4 mg of oligomycin were dissolved in 1 ml ethanol in a glass vial. This solution was divided and stored in glass vials at -20 °C.
- In the reported experiment 4 μl of 4 mg/ml Oligomycin were used (final concentration 10 μM).
- As shown in the figure, yeast cells did not respond to oligomycin and the respiration rate remained unchanged at the level of routine respiration.
- This has been observed by us also for the CEN.PK 113-7D strain in a similar protocol.
- A similar resistance to oligomycin of yeast has been recently reported also in:
- Leadsham JE, Gourlay CW. cAMP/PKA signaling balances respiratory activity with mitochondria dependent apoptosis via transcriptional regulation. BMC Cell Biol. 2010 Nov 25;11:92. doi: 10.1186/1471-2121-11-92. »Link
Popular Bioblast page
- Oligomycin has been accessed more than
- 15,000 times (2019-12-11)
- 10,000 times (2016-05-16)
- 5,000 times (2013-10-17)
- Oligomycin has been accessed more than