Kiss 2012 Abstract Bioblast
| Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C (2012) The negative impact of alpha-ketoglutarate dehydrogenase complex deficiency on matrix substrate-level phosphorylation. Mitochondr Physiol Network 17.12. |
Link: MiPNet17.12 Bioblast 2012 - Open Access
Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C (2012)
Event: Bioblast 2012
Objectives: Provision of succinyl-CoA by the alpha-ketoglutarate dehydrogenase complex (KGDHC) is essential for generation of matrix ATP (or GTP) by substrate-level phosphorylation catalyzed by succinyl-CoA ligase. A decline in KGDHC activity has been associated with neurodegeneration.
Methods: Mitochondrial phosphorylation was investigated in tissues of transgenic mice with deficiencies in KGDHC subunits.
Results: We demonstrate ATP consumption in respiration-impaired isolated and in situ neuronal somal mitochondria from transgenic mice with a deficiency of either dihydrolipoyl succinyltransferase (DLST) or dihydrolipoyl dehydrogenase (DLD) exhibiting a 20-48% decrease in KGDHC activity. Import of ATP into the matrix of mitochondria from transgenic mice was attributed to a shift in the reversal potential of the adenine nucleotide translocase towards more negative values due to diminished matrix substrate-level phosphorylation, causing the translocase to reverse prematurely. Immunoreactivity of all three subunits of succinyl-CoA ligase and maximal enzymatic activity were unaffected in transgenic mice as compared to wild-type littermates. Therefore, decreased matrix substrate-level phosphorylation was due to diminished provision of succinyl-CoA. These results were further corroborated by the finding that mitochondria from wild-type mice respiring on substrates supporting substrate-level phosphorylation exhibited ~30% higher ADP-ATP exchange rates compared to those obtained from DLST+/- or DLD+/- littermates.
Conclusions: We propose that KGDHC-associated pathologies are subserved by the inability of respiration-impaired mitochondria to rely on “in-house” mitochondrial ATP reserves.
• Keywords: Succinyl-CoA ligase, Adenine nucleotide translocase, F0-F1 ATP synthase, Reversal potential
• O2k-Network Lab: HU Budapest Chinopoulos C
Labels: MiParea: Respiration, Genetic knockout;overexpression Pathology: Aging;senescence Stress:Ischemia-reperfusion Organism: Mouse Tissue;cell: Nervous system, Liver Preparation: Intact cells, Isolated mitochondria, Enzyme Enzyme: Adenine nucleotide translocase, Complex V;ATP synthase, TCA cycle and matrix dehydrogenases Regulation: mt-Membrane potential, Substrate
HRR: Oxygraph-2k, O2k-Fluorometer
Affiliations and author contributions
Gergely Kiss (1), Csaba Konrad (1), Judit Doczi (1), Anatoly A Starkov (2), Hibiki Kawamata (2), Giovanni Manfredi (2), Steven F Zhang (2), Gary E Gibson (3), M Flint Beal (2), Vera Adam-Vizi (1), Christos Chinopoulos (1,2)
(1) Department of Medical Biochemistry, Semmelweis University, Budapest, 1094, Hungary; Email: [email protected]
(2) Weill Medical College Cornell University, New York, NY, 10021, USA
(3) Weill Cornell Medical College/Burke Medical Research Institute, White Plains, NY, 10605, USA
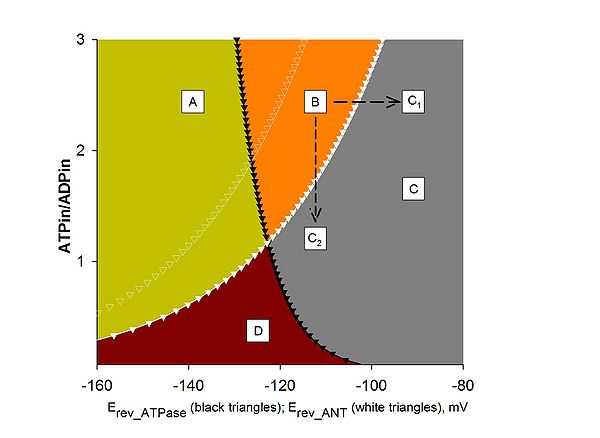
Figure 1
Computational estimation of the reversal potential of adenine nucleotide translocase (Erev_ANT) and reversal potential of F0-F1ATPase (Erev_ATPase). A: ATPase forward, ANT forward; B: ATP reverse, ANT forward; C, C1, C2: ATPase reverse, ANT reverse; D: ATPase forward, ANT reverse. Black solid triangles represent Erev_ATPase; white solid triangles represent Erev_ANT. Values were computed for [ATP]out = 1.2 mM, [ADP]out = 10 μM, Pin = 0.01 M, n = 3.7 (2.7 plus 1 for the electrogenic ATP4-/ADP3- exchange of the ANT), pHi = 7.38, and pHo = 7.25. White open triangles represent Erev_ANT values computed for [ATP]out = 1.4 mM, and all other parameters as above. Traces have been computed by Erev estimator; the software and instructions on how to use it can be downloaded here.